When my sister got her 23andme results, we sent them over to Doug McDonald. I was expecting something close to my results, but it was radically different:
This one is different it says 37% Druze, 4% Bushman or Pygmy, the rest North India. It is complicated enough that the program refuses to generate a spot on the map. The chromosome painting looks quite reasonable for that assignment.
I am including several plots .. these show just how odd this is.
Here are the PCA plots that Doug sent. My sister is shown by the crosshairs.
Think of this as two-dimensional projections of a multidimensional space and you’ll notice that my sister is not close to any of the reference groups.
You can see her 3-D position (“Test Person”) in the animation below (or by clicking on animation).
Her chromosome painting, a similar concept to 23andme’s ancestry painting, shows which chromosome segments are most like some population. As you can see, there are a few chromosomes that have almost no “South Asian” segments.
I was very surprised by my sister’s results, especially the 4% Bushman/Pygmy. I expected some East African admixture due to the Egyptian ancestry but no Pygmy. Also, I expected some (10-20%) Middle East contribution but Druze at 37% is just too high. So I asked Doug McDonald to redo my ancestry analysis with the new version of his software.
Here’s what he told me:
It says you are half North India, 3% Bushman or Pygmy, and the rest Iranian, OR 80% Sindhi, 2% Bushman or Pygmy, the rest being Bedouin.
The spot on the map is far SW Pakistan.
The Pygmy is clearly a mistake!
The Pygmy is definitely a mistake. Pygmies are a very distinctive population and because genetic diversity is very high in Africa, the continent of humanity’s origin, sometimes these reference populations can give weird results. These analyses basically try to fit your genetic data to reference populations’ data samples. That’s one reason why you see Sindhi or Pathan as a result for Punjabis because there are no Punjabis in the reference data of HapMap or HGDP.
Here are my PCA plots:







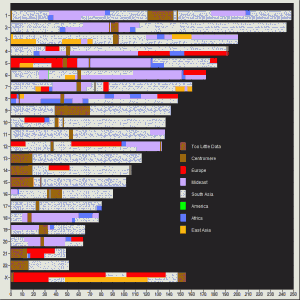



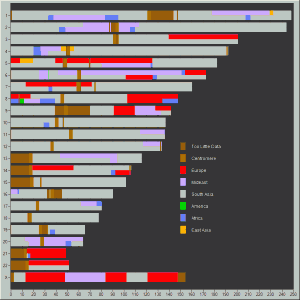
1 comment
Comments are closed.